In this tutorial, we will study the MySQL WEEKDAY() function. A week has seven days. Suppose you want to find the weekday number of any given date. This is where we use the MySQL WEEKDAY() function.
WEEKDAY() function is used to return a weekday number (called weekday index) for a given date. It treats Monday as the first day of the week. The weekday index starts at 0 which means the indexes for the weekdays are as follows –
- Monday = 0
- Tuesday = 1
- Wednesday = 2
- Thursday = 3
- Friday = 4
- Saturday = 5
- Sunday = 6
So if we pass a date to the WEEKDAY() function and it returns 2, that means that date is on a Wednesday. Let us now take a look at the syntax and some examples.
Syntax of MySQL WEEKDAY()
WEEKDAY(date)Code language: SQL (Structured Query Language) (sql)Where ‘date’ is the date/datetime value from which you want to extract the weekday index.
Examples of MySQL WEEKDAY()
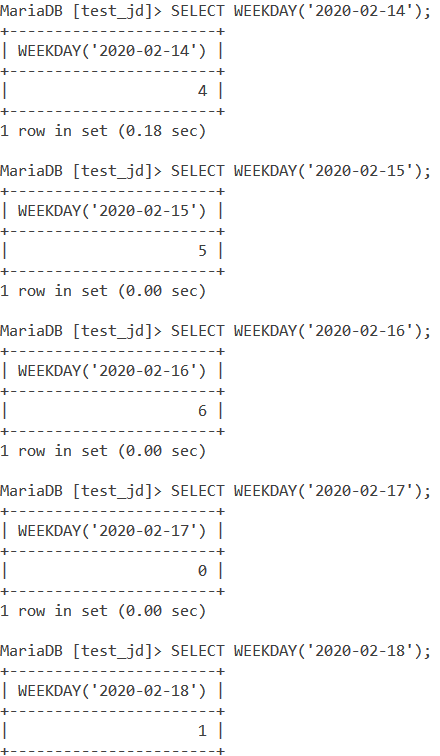
Let us take a look at some basic examples first. Let us find out the weekday index of the following dates using the WEEKDAY() function and the SELECT statement – 2020-02-14, 2020-02-15, 2020-02-16, 2020-02-17 and 2020-02-18.
SELECT WEEKDAY('2020-02-14');
SELECT WEEKDAY('2020-02-15');
SELECT WEEKDAY('2020-02-16');
SELECT WEEKDAY('2020-02-17');
SELECT WEEKDAY('2020-02-18'); Code language: SQL (Structured Query Language) (sql)And the output is –
MySQL Weekday() With Wrong Date Values
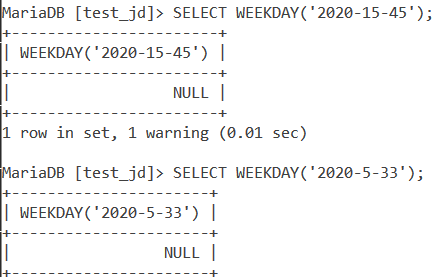
The WEEKDAY() function returns NULL if we pass it a wrong date value. A wrong date value is one in which the month value is greater than 12 or the day value is greater than 31. Let us see a couple of queries showing this example.
SELECT WEEKDAY('2020-15-45');
SELECT WEEKDAY('2020-5-33');Code language: SQL (Structured Query Language) (sql)And the output is –
WEEKDAY() With Datetime Values
As I mentioned in the syntax, we can pass datetime values as well to the MySQL Weekday() function. Let us see an example of this. Consider the below queries –
SELECT WEEKDAY('2014-12-31 00:01:00');
SELECT WEEKDAY('2020-02-01 20:51:10');Code language: SQL (Structured Query Language) (sql)And the output is –

WEEKDAY() and DAYNAME()
In a previous article on the DAYNAME() function, we saw that DAYNAME() returns the weekday name of the given date. WEEKDAY(), as you know, returns the weekday index number of the given date. Let us take a look at some queries to draw a correlation between these two functions. The queries are –
SELECT WEEKDAY('2020-02-14'), DAYNAME('2020-02-14');
SELECT WEEKDAY('2020-02-15'), DAYNAME('2020-02-15');
SELECT WEEKDAY('2020-02-16'), DAYNAME('2020-02-16');
SELECT WEEKDAY('2020-02-17'), DAYNAME('2020-02-17');
SELECT WEEKDAY('2020-02-18'), DAYNAME('2020-02-18'); Code language: SQL (Structured Query Language) (sql)And the output is –

As you can see, WEEKDAY() returns the index 4 when the DAYNAME() value is Friday and so on.
Using WEEKDAY() With Tables
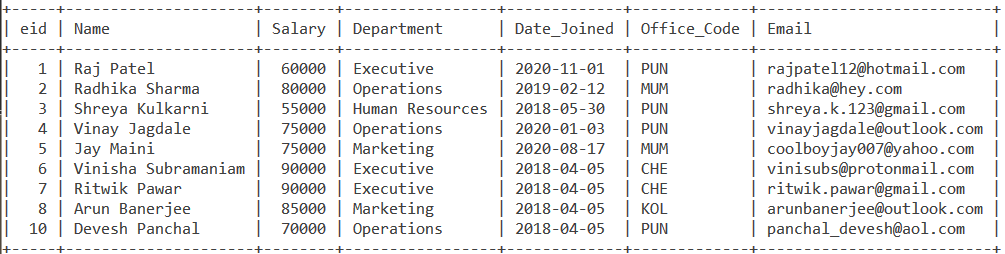
Consider the below Employee table.
Simple Example
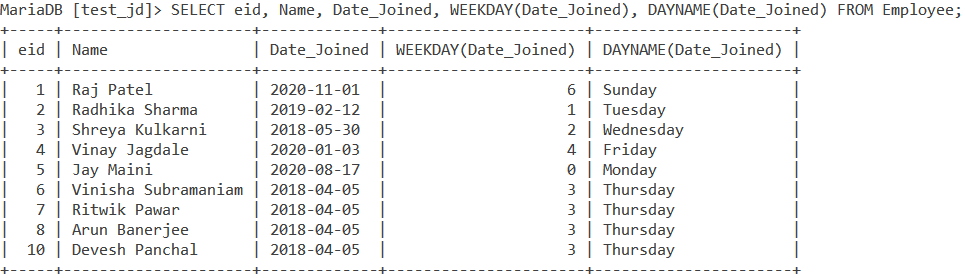
Building on that example in which we drew the correlation between WEEKDAY() and DAYNAME(), let us write a query that displays the employee id, Name of the employee, the date they joined and the weekday index and weekday name on which they joined. The query is –
SELECT eid, Name, Date_Joined, WEEKDAY(Date_Joined), DAYNAME(Date_Joined) FROM Employee; Code language: SQL (Structured Query Language) (sql)And we get the output as –
Using WHERE and IN with WEEKDAY()
Let us use the WHERE clause and the IN operator in a query along with the WEEKDAY() function. Let us write a query to find out which employees joined the company on weekends (they deserve an award for that), that is, Saturday or Sunday. The query for finding these hardworking employees is –
SELECT eid, Name FROM Employee WHERE WEEKDAY(Date_Joined) IN (5, 6);Code language: SQL (Structured Query Language) (sql)And the output is –

Conclusion
The WEEKDAY() function is widely used while retrieving the index number of the weekday from a date value. I would encourage you to practice different examples with different MySQL clauses.
References
- MySQL Official Documentation on
WEEKDAY().
